Error Analysis – SARS-CoV-2 variant detection
Bachelor thesis, completing the bachelor of Computer Science and Engineering at TU Delft.
Grade: 9.0
Research published as part of ‘Lineage abundance estimation for SARS-CoV-2 in wastewater using transcriptome quantification techniques. Baaijens, Jasmijn A.’ (https://doi.org/10.1186/s13059-022-02805-9).
Thesis abstract:
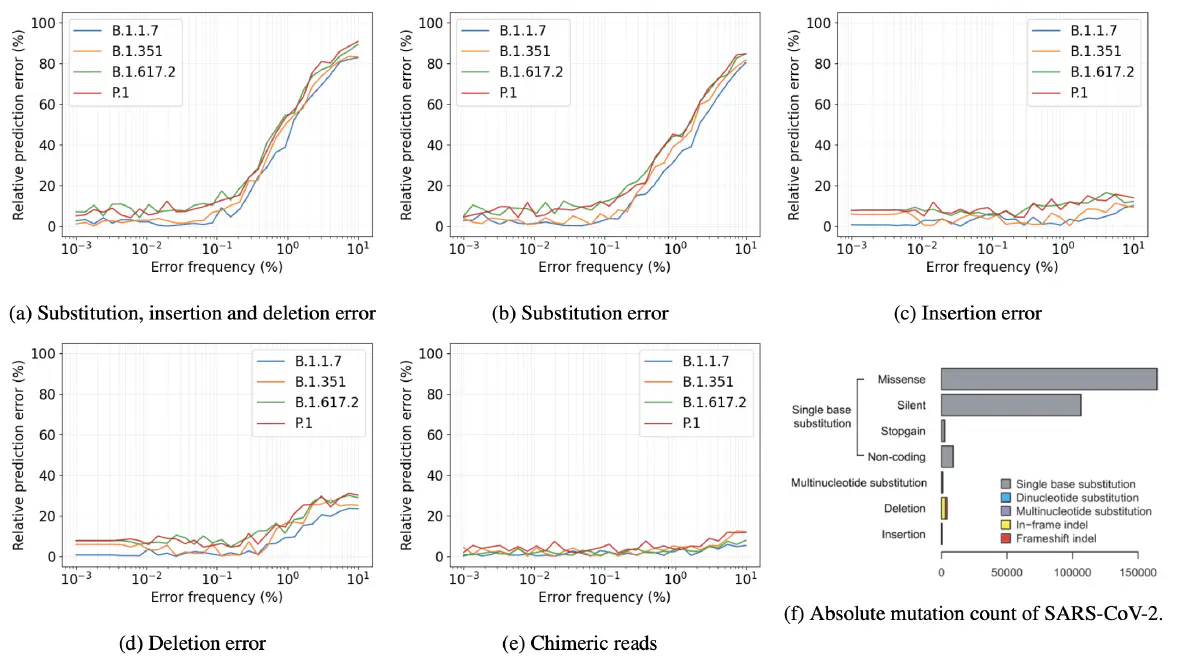
Since the start of the SARS-CoV-2 pandemic, the monitoring of SARS-CoV-2 by way of viral RNA sequencing of wastewater has proven to be an efficient and effective way of estimating COVID-19 cases in population groups. A recently developed pipeline also enables us to estimate SARS-CoV-2 variant abundance using viral samples from wastewater. This is done by repurposing an RNA-seq quantification algorithm to quantify reads, belonging to variants, from DNA-sequencing data. However, the impact of sequencing errors and contaminating viruses on this process is unknown. Here I show that, in simulated data, the credibility of the prediction results is dependent on the error rate of the sequencing machines used. I also show that contaminating the simulated dataset with certain human coronaviruses has a significant effect on prediction accuracy. However, most viruses currently found in wastewater have no effect. Furthermore, adding a reference genome for these human coronaviruses to the reference set removes any impact. The results demonstrate that it is important to assess the credibility of the pipeline on a case by case basis and to tailor the testing setup and reference set to this assessment.